The back-story of Pathomation is well-known: once upon a time, there was an organization that had many WSI scanners. All these scanners came with different pieces of software, which made it inconvenient for pathologists to operate digitally. After all, you don’t alternate between Microsoft Office and Apache OpenOffice depending on what kind of letter you want to write and which department you want to address it to.
Tadaaaah, ten years later Pathomation’s PMA.core tile server acts as a veritable Rosetta stone for a plethora of scanner vendors, imaging modalities, and storage capacities alike.

There’s a second story, too, however: Pathomation could not have been founded with a chance encounter of a couple of people with exactly the right background at the right time. As it happened, in 2012, the annual Digital Pathology Association in Baltimore (coinciding with a pass-through of hurricane Sandy) had a keynote speech by John Tomaszewski about the use of bioinformatics tools to (amongst others) determine Gleason scores on prostate slides. One of the co-founders of Pathomation was in the audience and thought “what a great idea…”.

Bioinformatics meets digital pathology
Pathomation reached out to the bioinformatics community early on. Within the ISCB and ECCB communities however, interest was initially low: only one or two talks (or even posters) at the Vienna, Berlin, and Dublin editions discussed anything even remotely related to microscopy. The few people that did operate in this intersection of both fields, expressed mostly frustration having to spend a seemingly outrageous amount of time just learning how to extract the pixels from the raw image data files.
A simple observation emerged: bioinformatics could contribute a lot more to digital pathology, if only we could make it easier to port the data back and forth between the two communities. Say: A universal honest broker for whole slide imaging.
But having such software (in the form of its CE-IVD certified PMA.core tile server and freeware “starter kit” PMA.start) is not enough. We still had to get the word out.
So Pathomation next set out on its own and started organizing its own digital / computational pathology workshops. Coinciding with the European-based bi-annual ECCB events in The Hague (The Netherlands) and Athens (Greece) The proceedings of these are still available online.
A maturing relationship
Fast forward to 2022 and things are a bit different.
As Pathomation has forged on as a middleware provider, so has bioinformatics gradually started to contribute to digital pathology. As we already hypothesized 10 years ago: the datasets in pathology (and by extension: microscopy, histology, time lapse data, MSI etc) are too interesting not to be explored with tools that have already contributed so much to other fields like genetics and medicine.
When you go to OUP’s leading Bioinformatics journal and search for “pathology”, more than 10,000 hits spring up.
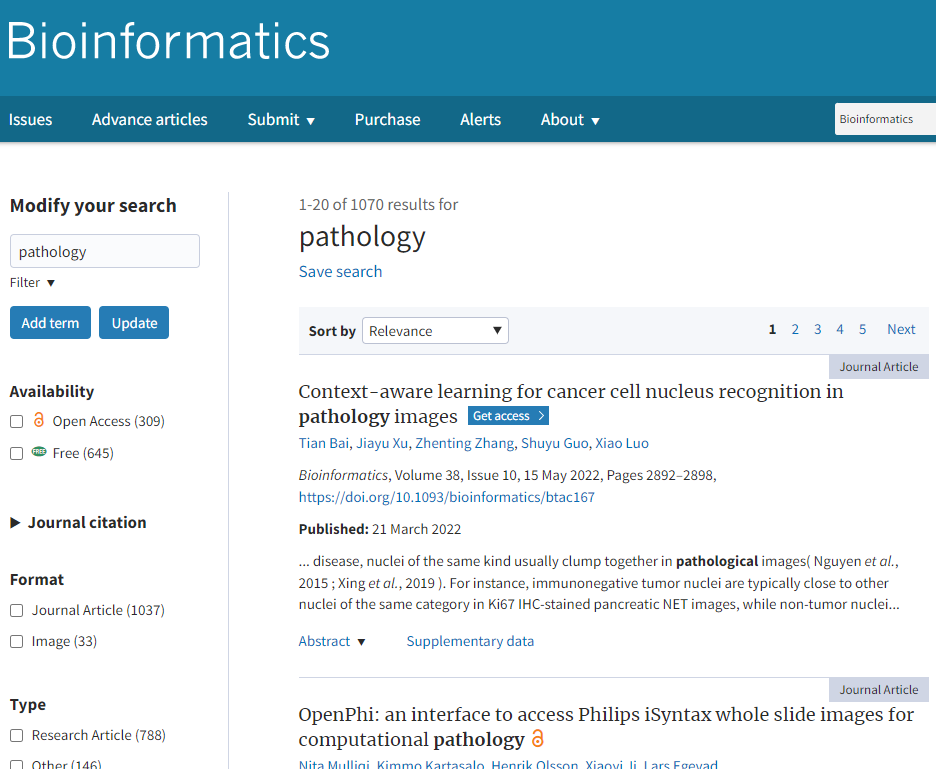
Correspondingly, a search for “bioinformatics” in the Journal of Pathology Informatics (JPI) yields significantly fewer results. That’s not unexpected, as oftentimes “bioinformatics” wouldn’t be mentioned as a wholesale term, but one would rather reference a particular protocol, method or technique instead. Chubby Peewee, anyone?

The relationship between digital pathology and bioinformatics is clearly well established and maintained.
Building bridges
Occasionally we’re asked what we offer in the field of image analysis (IA). We’ve always maintained our stance that we don’t want to get into this directly ourselves by means of offering an alternative product (or software component) to well established packages like ImageJ, QuPath (open source) or Visiopharm, HALO, or Definiens (commercial).
Instead we’ve pursued our mission of becoming the one true agnostic broker for digital pathology. This means that we look at all of these (and more) and determine how we can best contribute to help people transfer information back-and-forth between different environments.

SDKs
Many in silico experiments start of as scripts. In bioinformatics, one of the first extensively supported scripting languages was Perl, in the form of the BioPerl framework. It still exists and is in use today, but (at the risk of alienating Perl-aficionados; we mean no offense) has been surpassed in popularity by Python (and BioPython).
Looking at the success of (bio)python, the first language we decided to support in the form of providing an SDK for it was Python: our PMA.python library is included in the Python Package Index (PyPI), and very easy to install through an interactive framework like Jupyter or Anaconda.
Several articles on this blog tackle digital pathology challenges by means of our own PMA.python library, and can be used as an inspiration for your own efforts:
All our SDKs are available as open source through GitHub, and since PMA.python, we’ve also added on PMA.java and PMA.php.
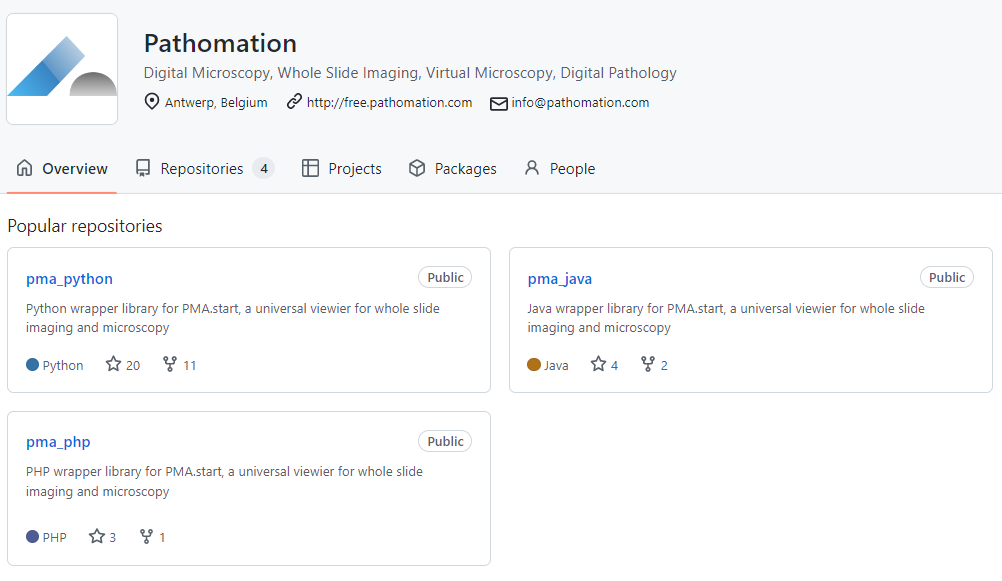
We have a development portal with sections on each programming language that we support, and we use the respective “best practice” mechanisms in each to provide exhaustive documentation on provided functions and methods as well.
Plugins
The plugin concept is a powerful one: many companies that build broadly used software have realized that they can’t themselves foresee all uses and provide a way for third-party providers to extend the software themselves through a plugin mechanism. In fact, software has a better chance of becoming broadly used when it offers this kind of functionality from the start. It’s called “the network effect”, and the network always wins.
Not everybody wants (or needs) to start from the ground up. There are many environments that can serve as a starting point for image analysis. Two specific ones related to digital pathology are ImageJ and QuPath. As we needed a proving ground for our own PMA.java SDK anyway, we decided to work towards these two environments. Today, we have plugins available for both.
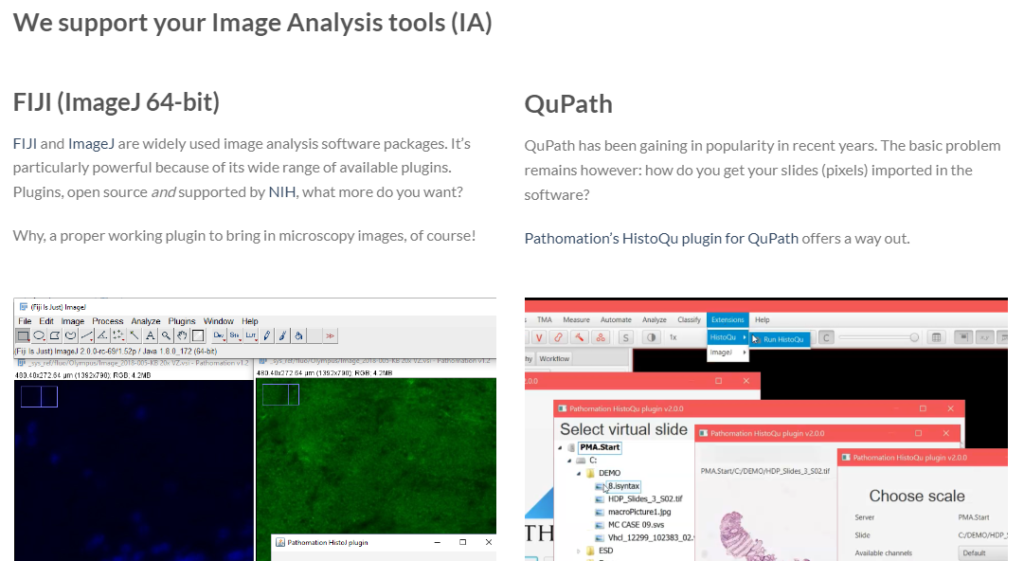
Our plugins are available from our website, free of charge. You can download our plugins for image analysis, which are bioinformatics related. For more mundane applications, we also offer plugins to content management systems (CMS) and learning management systems (LMS).
The PMANN interface
If there is no plugin architecture available, oftentimes an intermediary file can be used to exchange data. We apply this technique ourselves in PMA.studio, where a virtual tray can persist over times by exporting it to and importing it back from a CSV file as needed.
So it is with commercial providers. Visiopharm has its own (binary) MLD file format to store annotations in, and Indica Labs’ HALO uses an XML-based file.
What you typically want to do, is run an algorithm repeatedly, with slightly different parameters. This results then in different datasets, that are subject to comparison and often even manual inspection or curation.
The PathoMation ANNotation interface assists in this: As the analytical environments are oftentimes unsuited for curators to work with (both in terms of complexity as well as monetary cost), a slide in PMA.core can be instructed to look for external annotations in various locations. You can have slide.svs associated with algo1.mld, algo2.mld, and algo3.mld. You can interpret the overlaying datalayers computationally, or you can visualize them in an environment like PMA.studio.
What’s more: PMANN allows you to overlay data from multiple environments simultaneously. So you can do parameter optimization in one environment, but you can also run an algorithm across different environments and see which environment (or method like deep learning or random forest) performs the best.
Don’t reinvent the wheel
Bioinformatics is a much more mature field than digital pathology, with a much broader community. At Pathomation, we strongly believe that many techniques developed are transferable. Data-layers and data-sets are equally apt to enrich each other. Genetics and sequencing offer resolution, but tissue can add structure as an additional source of information to an experiment outcome, and help with interpretation. The work of Ina Koch that we referenced earlier is a typical example of this. Another great example is the TCGA dataset, which has been incorporating whole slide images as well as sequencing data for a couple of years now.
At Pathomation, we aim to offer connectivity. Connectivity to different slide scanner vendors (by supporting as many file formats as possible), but also by allowing different software tools to exchange data in a standard manner. To this end, Pathomation also recently signed up for the Empaia initiative.

We strongly believe that as a physician or researcher alike, you should be able to focus on what you want to do, rather than how you want to do it. We build middleware, SDKs, and plugins to help you do exactly that.
Do you have experience with coupling bioinformatics and digital pathology? Have you used our software already in your own research? Let us know, and you may find yourself the subject of a follow-up blog post.
