Now that both PHP and Java have methods for embedded slide visualization, we can’t leave Python out. Originally we didn’t think there would be much need for this, but it’s at least confusing to have certain methods available in one version of our SDK, while not in others.
In addition, interactive visualization is definitely a thing in Python; just have a look at what you can do with Bokeh. Ideally and ultimately, we’d like to add digital pathology capabilities to an already existing framework like Bokeh, but in this blog post we’ll just explore how you can embed a slide into your IPython code as is.
As PMA.python is not a standard library, it bears to start your notebooks with the necessary detection code for the library. If it doesn’t work, it’s bad manners to leave your users in the dark, so we’ll provide some pointers on what needs to be done, too:
try:
from pma_python import core
print("PMA.python loaded: " + core.__version__)
except ImportError:
print("PMA.python not found")
print("You don't have the PIP PMA.python package installed.\n"
+ "Please obtain the library through 'python -m pip install pma_python'")
If all goes well, you’ll see something like this:
Once you’re assured the PMA.python library is good to go, you should probably verify that you can connect to your PMA.core instance (which can be PMA.start, too, of course; just leave the username and password out in that case):
server = "http://yourserverhere/pma.core/"
user = "your_username"
pwd = "your_password"
slide = "rootdir/subdir/test.scn"
session = core.connect(server, user, pwd)
if (session):
print("Successfully connected to " + server + ": " + session)
else:
print("Unable to connect to PMA.core " + server + ". Did you specify the right credentials?")
If all goes well, you should get a message that reads like this:
Successfully connected to http://yourserverhere/pma.coreFinally, the visualization part. Note that Pathomation provides a complete front-end Javascript-framework for digital pathology. In order to bring these capabilities into (I)Python then, it sufficient to write some encapsulation code around this basic demonstration code:
def show_slide(server, session, slide):
try:
from IPython.core.display import display, HTML
except ImportError:
print("Unable to render slide inline. Make sure that you are running this code within IPython")
return
render = """
<script src='""" + server + """scripts/pma.ui/pma.ui.view.min.js' type="text/javascript"></script>
<div id="viewer" style="height: 500px;"></div>
<script type="text/javascript">
// initialize the viewport
var viewport = new PMA.UI.View.Viewport({
caller: "Jupyter",
element: "#viewer",
image: '""" + slide + """',
serverUrls: ['"""+ server + """'],
sessionID: '""" + session + """'
},
function () {
console.log("Success!");
},
function () {
console.log("Error! Check the console for details.");
});
</script>"""
display(HTML(render))
Our method is a bit more bulky than strictly needed; it’s robust in this sense that it makes sure that it is actually running in an IPython environment like Anaconda, and will also provide output to the JavaScript console in case the slide can load for some reason.
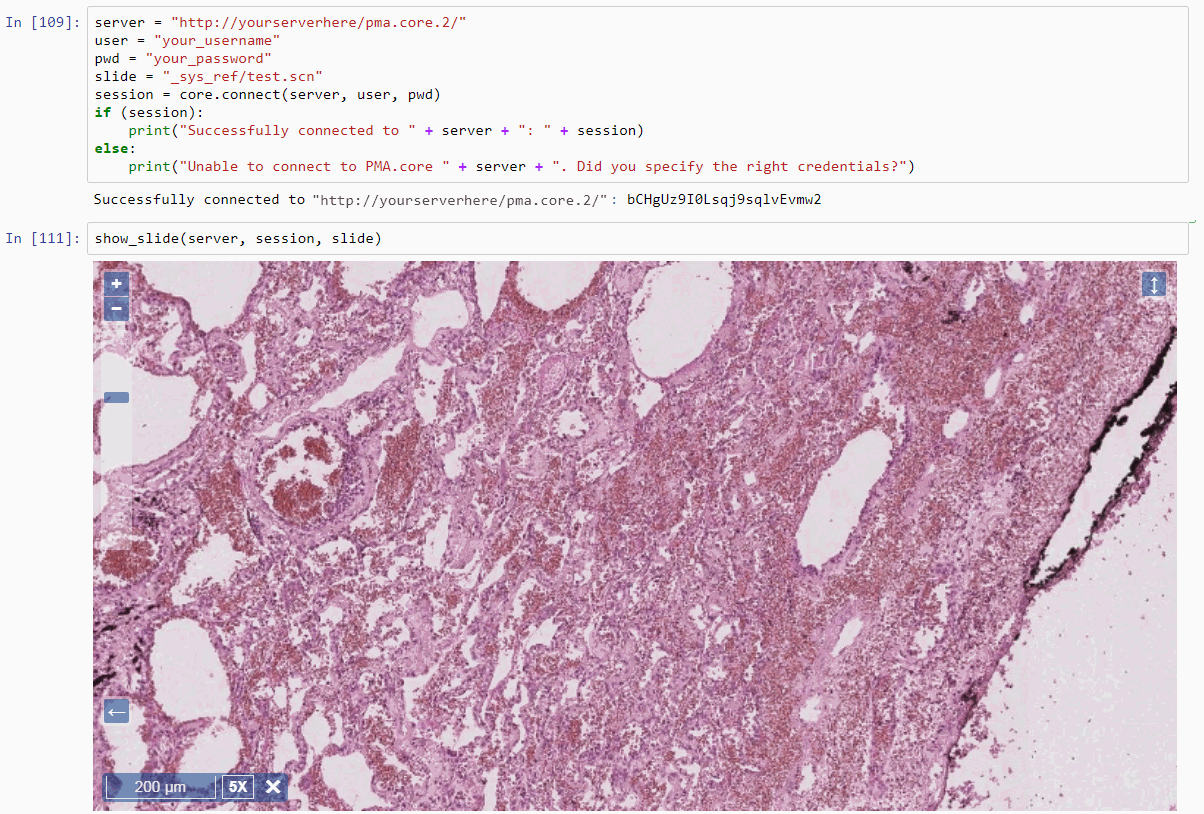
Rendering a slide inline within a Python / Jupyter notebook is now trivial (just make sure you ran the cell in which you define the above method, before invoking the following piece of code):
show_slide(server, session, slide)
The result look like this:
There is never an excuse not to use exploratory data analysis to get initial insights in your data. As switching environments, browsers, screens… can be tedious, and notebooks are meant to encapsulate complete experiments, interactive visualization of select whole slide images may just be one more thing you want to include.
The .ipynb file can be downloaded here and used as a starting point in your own work.
By studying the PMA.UI framework, you can learn more about how to further modify and customize your interactive views.
Now, anybody out there who wants to pick up our Bokeh challenge?